Wiki Your Genes
Kevin Kelly
February 26, 2008
I am taking a crash course in genetic literacy by having some of my genes sequenced by the two major genetic sequencing services, 23andMe and deCode. I am still in the process of comparing the two sets of results to see which vendor is better, but while coming up to speed in this new realm, David Ewing Duncan, another self-experimenter, turned me onto a very cool site: SNPedia.
SNPedia is a wiki for personal genomic raw data, which come in SNPs, or in my usage, snips.
Snips are the current desired unit in personal genomics, like pixels in digital photography. For now, more snips is better. A snip is a particular part of the gene that researchers have noticed will vary between individuals (most of the gene does not vary). These single-nucleotide polymorphism (SNP) variations are the tiny spots on your chromosome that are actually sequenced and reported back to individuals. Each snip position has a unique number, and some of these snips such as rs1815739 (good sprinter) or rs795174 (green eye color) indicate particular traits.
On SNPedia you can paste in a snip number — from the results of your DNA sequencing — and find out what is currently known about it. Or conversely you can enter a trait or disease and see if there is a snip tagged to it. As information is gleaned from medical journals, wikians add it to the SNPedia. Usually very quickly.
A typical entry will look like this:
rs7495174 is located in intron 1 of the OCA2 gene. The (A) allele (in dbSNP orientation) is associated with blue or green eye color in Caucasians. [PMID 17236130].
This SNP is 1 of 3 SNPs defining a haplotype that has been studied for association with eye color. The full details on the correspondence between the haplotype and eye color can be found on the OCA2 page.
In theory this is what the news personal genomics sites of 23andMe and deCode are supposed to be doing. Only with slick user-friendly designs. They do offer this information, but in their effort to filter this large deluge of data, they both are selecting certain snips as being more important/interesting, and hiding the rest in the page pages of their sites. To surf the ocean of data beyond these selected traits, diseases, or snips is cumbersome. And of course, it costs $1,000 to enter the door — the price of getting your DNA sequenced at either place.
Here is how I see the nascent field of personal genomic testing shaping up. The retailers are 23andMe and deCode. They don’t sequence genes. They outsource that specialized job to microarray manufacturers, while the retailers sell the website interface, and supporting information to consumers. There are only two main manufacturers of the large scale mircoarray chip which is used to provide up to 1 million SNIPS. One is Affymetrix, and the other in the Illumina. Affymetrix and Illumina will sell their microarrays to anyone, although they currently don’t sell to individuals. Affymetrix lists the cost of a 500,000 SNIP array chip at $250 today. You need their specialized machines and software to read the chip.
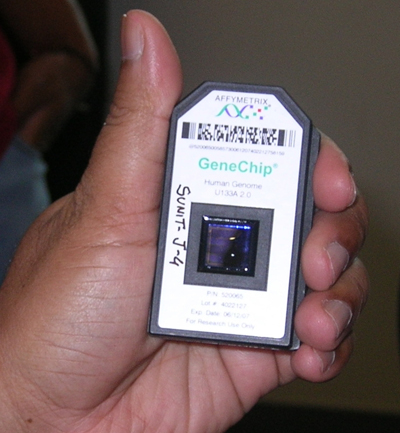

If a third-party vendor were to start selling the naked chip’s data for a small fee above its costs, it would be possible to do a large personal sequence using one of these tests and managing your data using open-source wiki technology like SNPedia. Hard-core recreational genomist could probably do a better job than either 23andMe or Decode are doing right now.
This is close to a DIY kit for geneboys. With some mashing of websites, you could get more info in, faster, more personalized to you. If you are already doing this, write me.